A massive genomic study in Sardinia has discovered a protective variant against malaria in the CCND3 gene. This mutation, a product of evolutionary adaptation when the disease was endemic, produces larger red blood cells that suffocate the parasite. The research, published in Nature, not only explains a unique population trait but also offers a molecular template for future drugs. From visual epidemiology, this finding is a perfect case for modeling in 3D the intersection between genetics, history, and geography of a disease. 🧬
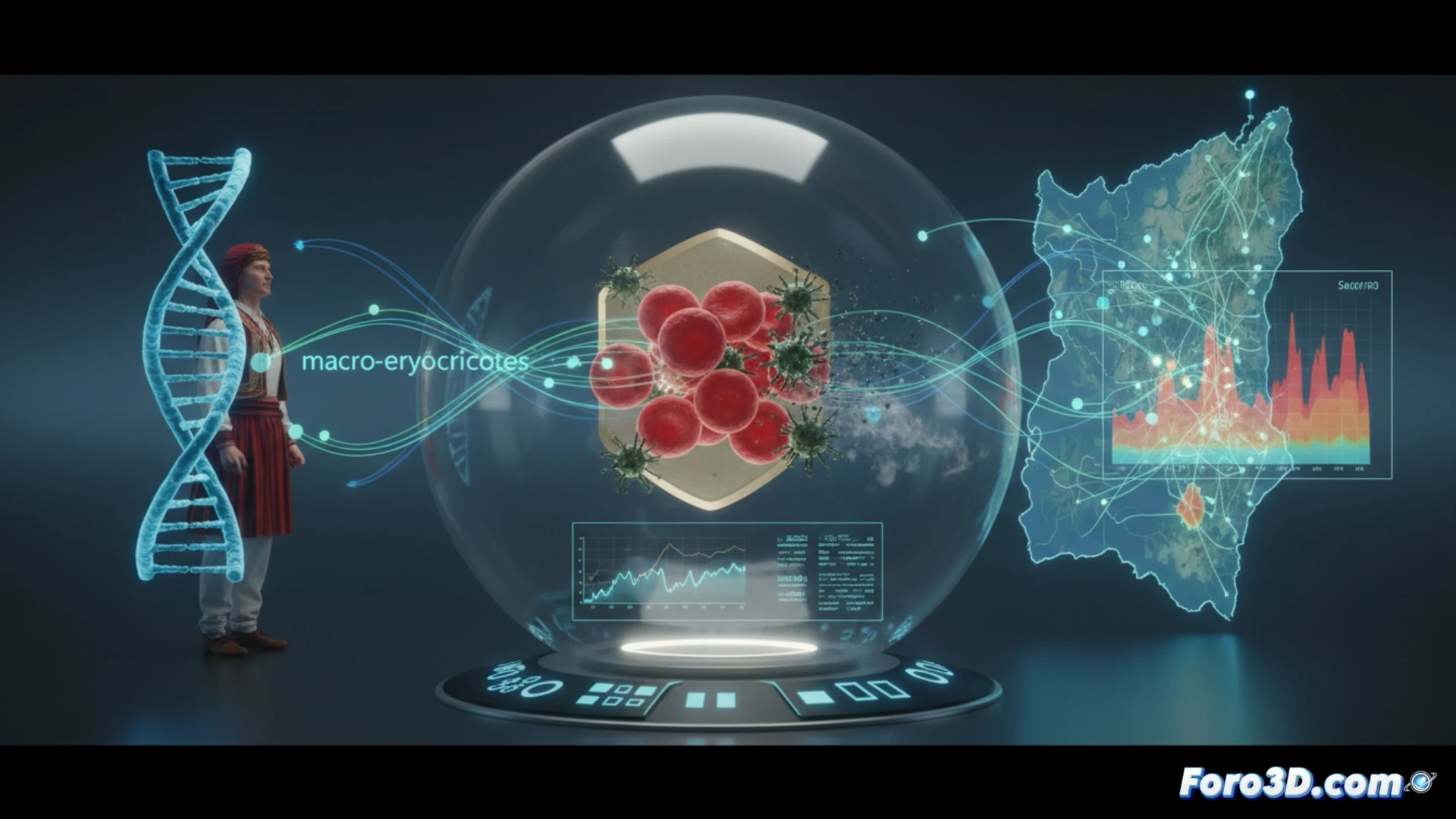
3D Visualization of a Genetic Mechanism and Its Geographic Distribution 🗺️
The power of this study for our niche lies in its ability to be visualized. An interactive model with two main layers is proposed. First, a 3D map of Sardinia showing the frequency of the genetic variant in different districts, overlaid on historical layers of malaria incidence. Second, a cellular-scale animation detailing the mechanism: how the variant in CCND3 alters red blood cell maturation, increasing their volume and modifying their internal environment, leading to the death of Plasmodium. This dual visualization connects population phenomenon and molecular mechanism.
The Genetic Trail of Epidemics and the Future of Public Health 💊
This case demonstrates how the human genome is an archive of our epidemiological battles. Visualizing this data makes tangible the footprint that a disease, now eradicated in a territory, leaves on the biology of its inhabitants. Additionally, modeling this natural defense mechanism in 3D provides an invaluable tool for the rational design of therapies that mimic it. Thus, visual epidemiology becomes a bridge between adaptive past and future pharmaceutical innovation.
How would you create an interactive map that shows data by age groups and sex?